Another paper suggests an early peopling of America: Human Y chromosome sequences from Q Haplogroup reveal a South American settlement pre-18,000 years ago and a profound genomic impact during the Younger Dryas, Paula B. Paz Sepúlveda et al. ,Andrea Constanza Mayordomo,Camila Sala,Ezequiel Jorge Sosa,Jonathan Javier Zaiat,Mariela Cuello,Marisol Schwab,Daniela Rodríguez Golpe,Published: August 17, 2022 https://doi.org/10.1371/journal.pone.0271971.
This is one of a growing trend that supports an early arrival of human beings to America, in this case before 18,000 years BP.
The abstract states: "The present is the first genomic study of Q Haplogroup in which current knowledge on Q-M848 sub-lineages is contrasted with the historical, archaeological and linguistic data available. The divergence times, spatial structure and the SNPs found here as novel for Q-Z780, a less frequent sub-haplogroup autochthonous of the Americas, provide genetic support for a South American settlement before 18,000 years ago.
I foung it intersting that Argentine men of Native American origin shared a sub-linage with men from Sri Lanka! : "One of the new Argentine samples sequenced in this study, from San Juan (RUTBE), is presented as a sub-lineage of Q-F4674 along with 2 other Sri Lankan samples from the databases (S5 Fig and S1 Table). In turn, San Juan’s sample shares Q-Z36057 with 1 of Sri Lanka’s individuals; the age of this lineage has not been yet estimated. The occurrence of Q-F4674 in the Americas has recently been found and published by our group".
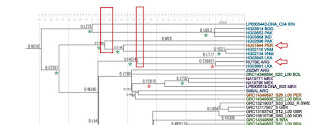
Below is the treee, the green asterisk marks dated lineages, notice how old this one is! and the Peruvian one almost 24 Ky old. But the paper didn't analyze it.
Patagonian Monsters - Cryptozoology, Myths & legends in Patagonia Copyright 2009-2022 by Austin Whittall ©